Collapsed variational inference for non-parametric Bayesian mixture models
Source:R/cvi_npmm.R
cvi_npmm.RdCollapsed variational inference for non-parametric Bayesian mixture models
Usage
cvi_npmm(
X,
variational_params,
prior_shape_alpha,
prior_rate_alpha,
post_shape_alpha,
post_rate_alpha,
prior_mean_eta,
post_mean_eta,
log_prob_matrix = NULL,
maxit = 100,
n_inits = 5,
Seed = NULL,
parallel = FALSE,
covariance_type = "full",
fixed_variance = FALSE,
cluster_specific_covariance = TRUE,
variance_prior_type = c("IW", "decomposed", "sparse", "off-diagonal normal"),
...
)Arguments
- X
input data as a matrix
- variational_params
number of clusters in the variational distribution
- prior_shape_alpha
shape parameter of Gamma prior for the DP concentration parameter alpha. Default is 0.001
- prior_rate_alpha
rate parameter of Gamma prior for the DP concentration parameter alpha. Default is 0.001
- post_shape_alpha
initial value for posterior update of shape parameter for alpha. Default is 0.001
- post_rate_alpha
initial value for posterior update of ratee parameter for alpha. Default is 0.001
- prior_mean_eta
mean vector of MVN prior for the DP mean parameters. Default is zero vector
- post_mean_eta
initial value of posterior update for the DP mean parameter
- log_prob_matrix
logarithm of cluster allocation probability matrix. Default is NULL
- maxit
maximum number of iterations. Default is 100
- n_inits
Number of random initialisations if log_prob_matrix and other case-specific hyperparameters are NULL. Default is 5
- Seed
Seeds for random initialisation; either a vector of n_inits integers or NULL. Default is NULL.
- parallel
Logical input for parallelisation. Default is FALSE
- covariance_type
covariance matrix is considered diagonal or full. Default is 'full'
- fixed_variance
covariance matrix of the data is considered known (fixed) or unknown. Default is FALSE
- cluster_specific_covariance
covariance matrix is specific to a cluster allocation or it is same over all cluster choices. Default is TRUE
- variance_prior_type
For unknown and full covariance matrix, choice of matrix prior is either Inverse-Wishart ('IW') or Cholesky-decomposed ('decomposed'). For unknown, full and cluster-specific covariance matrix, choice of matrix prior is either Inverse-Wishart ('IW'), element-wise Gamma and Laplace distributed ('sparse') or element-wise Gamma and Normal distributed ('off-diagonal normal')
- ...
additional parameters, further details given below
Value
[vimixr()] returns a list with the following elements:
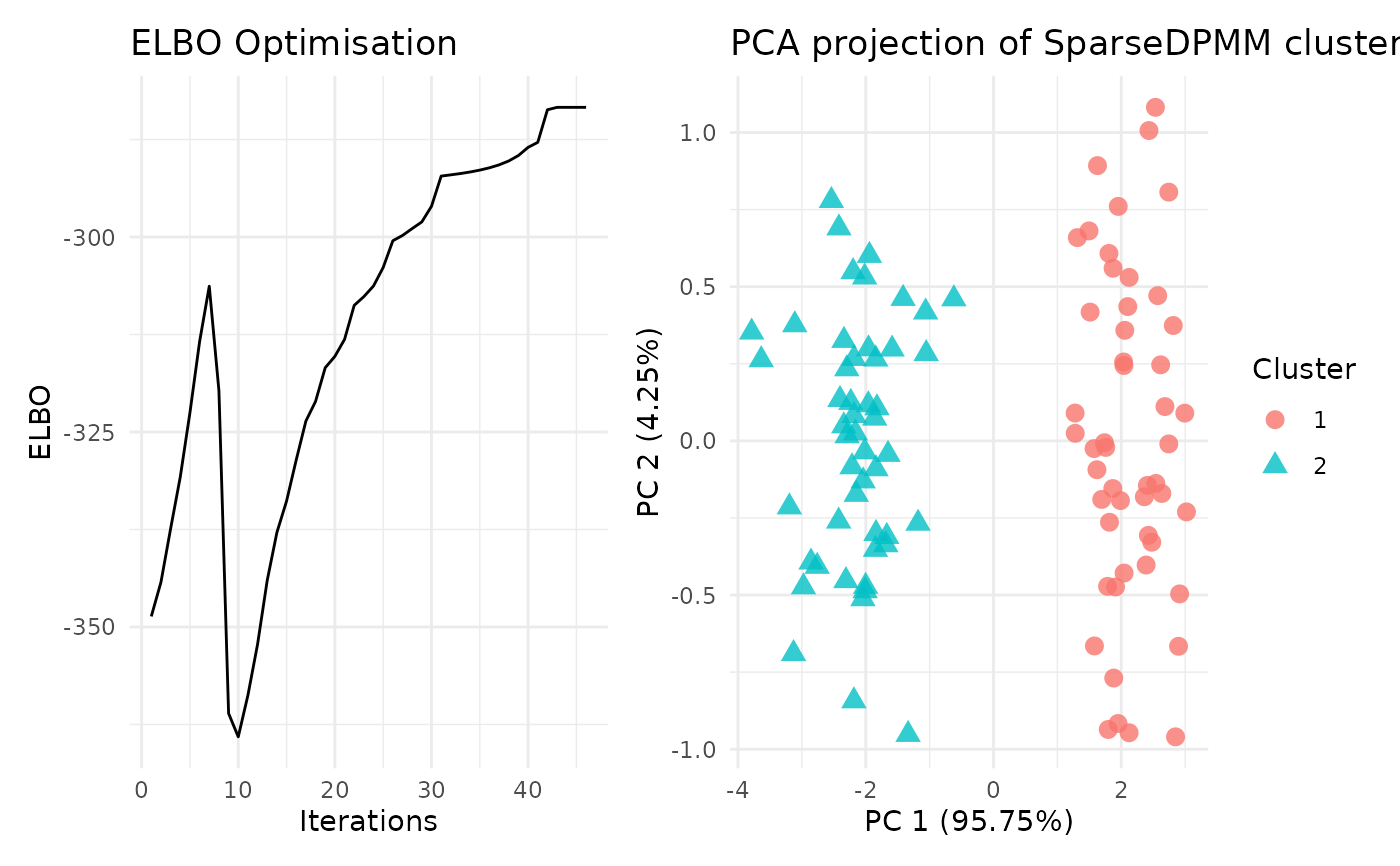
alpha: posterior DP concentration parameterCluster number: number of clusters from posterior probability allocation matrixCluster Proportion: cluster proportions from posterior probability allocation matrixlog Probability matrix: log of posterior probability allocation matrixELBO: Optimisation of the ELBO functionIterations: Number of iterations required for convergencePCA_viz: A PCA[ggplot2]plot to visualize the clustering of data based on cluster labelsELBO_viz: A line[ggplot2]plot to visualize the ELBO optimisation
Details
The following models are supported in vimixr, listing their
required input arguments in ... when calling cvi_npmm():
Known covariance
diagonal covariance We need the following additional arguments:
cov_data: a non-negative diagonal matrix, representing the covariance of the dataprior_precision_scalar_eta: a non-negative scalar, representing the precision prior for the DP mean parameterspost_precision_scalar_eta: initial value for the posterior update of precision for the DP mean parameters
full covariance We need the following additional arguments:
cov_data: a positive definite matrix, representing the covariance of the dataprior_cov_eta: a positive definite matrix, representing the covariance prior for the DP mean parameterspost_cov_eta: initial value for the posterior update of covariance for the DP mean parameters
Unknown covariance (Global)
diagonal covariance We need the following additional arguments:
prior_shape_scalar_cov: a non-negative scalar, representing the shape parameter of Gamma prior for the precisionprior_rate_scalar_cov: a non-negative scalar, representing the rate parameter of Gamma prior for the precisionpost_shape_scalar_cov: initial value for posterior update of precision shape parameterpost_rate_scalar_cov: initial value for posterior update of precision rate parameterprior_precision_scalar_eta: a non-negative scalar, representing the precision prior for the DP mean parameterspost_precision_scalar_eta: initial value for the posterior update of precision for the DP mean parameters
Inverse-Wishart We need the following additional arguments:
prior_df_cov: a scalar as the degree of freedom parameter of the Inverse-Wishart prior, Default value D+2prior_scale_cov: positive-definite matrix as the scale parameter of the Inverse-Wishart priorpost_df_cov: initial value for the posterior update of degree of freedompost_scale_cov: initial value for the posterior update of scale matrixprior_cov_eta: a positive definite matrix, representing the covariance prior for the DP mean parameterspost_cov_eta: initial value for the posterior update of covariance for the DP mean parameters
Cholesky-decomposition We need the following additional arguments:
prior_shape_diag_decomp: a non-negative scalar as the shape parameter of Gamma prior for diagonal elements of the Cholesly-decomposed matrixprior_rate_diag_decomp: a non-negative scalar as the rate parameter of Gamma prior for diagonal elements of the Cholesly-decomposed matrixprior_mean_offdiag_decomp: a scalar as the mean parameter of Normal prior for off-diagonal elements of the Cholesly-decomposed matrixprior_var_offdiag_decomp: a non-negative scalar as the variance parameter of Normal prior for off-diagonal elements of the Cholesly-decomposed matrixpost_shape_diag_decomp: initial value for posterior update of the shape parameter for diagonal elementspost_rate_diag_decomp: initial value for posterior update of the rate parameter for diagonal elementspost_mean_offdiag_decomp: initial value for posterior update of the mean parameter for off-diagonal elementspost_var_offdiag_decomp: initial value for posterior update of the variance parameter for off-diagonal elementsprior_cov_eta: a positive definite matrix, representing the covariance prior for the DP mean parameterspost_cov_eta: initial value for the posterior update of covariance for the DP mean parameters
Unknown covariance (cluster-specific)
Inverse Wishart We need the following additional arguments:
prior_df_cs_cov: a vector representing degree of freedom parameters for each cluster-specific Inverse-Wishart priorprior_scale_cs_cov: an array of positive-definite matrices representing scale matrix parameters for each cluster-specific Inverse-Wishart priorpost_df_cs_cov: initial value for posterior update of the degree of freedom parameterspost_scale_cs_cov: initial value for posterior update of the scale matrix parametersscaling_cov_eta: a non-negative scaling factor for covariance matrix of the DP mean parameters
Element-wise Gamma and Laplace prior We need the following additional arguments:
prior_shape_d_cs_cov: a non-negative vector as shape parameters for cluster-specific Gamma priors of the diagonal elementsprior_rate_d_cs_cov: a non-negative matrix as rate parameter for cluster-specific Gamma prior of the diagonal elementsprior_var_offd_cs_cov: a non-negative vector as variance parameter for cluster-specific Laplace priors of the off-diagonal elementspost_shape_d_cs_cov: initial value for posterior update of the diagonal shape parameterspost_rate_d_cs_cov: initial value for posterior update of the diagonal rate parameterspost_var_offd_cs_cov: initial value for posterior update of the off-diagonal variance parametersscaling_cov_eta: a non-negative scaling factor for covariance matrix of the DP mean parameters
Element-wise Gamma and Normal prior We need the following additional arguments:
prior_shape_d_cs_cov: a non-negative vector as shape parameters for cluster-specific Gamma priors of the diagonal elementsprior_rate_d_cs_cov: a non-negative matrix as rate parameter for cluster-specific Gamma prior of the diagonal elementsprior_var_offd_cs_cov: a non-negative scalar as variance parameter for cluster-specific Normal priors of the off-diagonal elementspost_shape_d_cs_cov: initial value for posterior update of the diagonal shape parameterspost_rate_d_cs_cov: initial value for posterior update of the diagonal rate parameterspost_mean_offd_cs_cov: initial value for posterior update of the off-diagonal mean parametersscaling_cov_eta: a non-negative scaling factor for covariance matrix of the DP mean parameters
Examples
X <- rbind(matrix(rnorm(100, m=0, sd=0.5), ncol=2),
matrix(rnorm(100, m=3, sd=0.5), ncol=2))
#for fixed-diagonal
res <- cvi_npmm(X, variational_params = 20, prior_shape_alpha = 0.001,
prior_rate_alpha = 0.001, post_shape_alpha = 0.001,
post_rate_alpha = 0.001, prior_mean_eta = matrix(0, 1, ncol(X)),
post_mean_eta = matrix(0.001, 20, ncol(X)),
log_prob_matrix = t(apply(matrix(-3, nrow(X), 20), 1,
function(x){x/sum(x)})), maxit = 100,
fixed_variance = TRUE, covariance_type = "diagonal",
prior_precision_scalar_eta = 0.001,
post_precision_scalar_eta = matrix(0.001, 20, 1),
cov_data = diag(ncol(X)))
#> outer loop: 1
#> -8.53924660769159-303.267632794874-175.811503063005-183.787706640935322.757259374774
#> outer loop: 2
#> -8.46140365758184-296.222923671738-175.979095995088-183.787706640935320.197518617843
#> outer loop: 3
#> -8.37788985971017-287.665654900053-176.963594667348-183.787706640935319.399929438151
#> outer loop: 4
#> -8.28767992870113-277.60102967719-177.040044575951-183.787706640935315.958870806487
#> outer loop: 5
#> -8.19057851715466-266.649475303788-178.016092022311-183.787706640935314.123947706829
#> outer loop: 6
#> -8.08599066456686-255.406903008855-179.078150029617-183.787706640935312.943910579916
#> outer loop: 7
#> -7.97343197786236-245.500391014289-180.266475916973-183.787706640935311.143136760766
#> outer loop: 8
#> -7.97452525802575-247.100211712943-181.121809063026-183.787706640935299.588154174956
#> outer loop: 9
#> -7.97219814739761-245.63545663705-181.952223813191-183.787706640935257.737569894688
#> outer loop: 10
#> -7.97120269168417-233.643645099042-182.061744159335-183.787706640935243.140238965843
#> outer loop: 11
#> -7.71360416439444-223.25796124581-182.956382837516-183.787706640935238.896325006473
#> outer loop: 12
#> -7.56188514912352-213.142612435777-183.017298142255-183.787706640935235.163625339611
#> outer loop: 13
#> -7.56007539267214-204.176185208136-184.95847006026-183.787706640935236.374187776412
#> outer loop: 14
#> -7.38862263841365-195.828749282045-185.730110698349-183.787706640935234.743296706177
#> outer loop: 15
#> -7.18994432479329-187.888140634882-185.967344196433-183.787706640935230.754606468997
#> outer loop: 16
#> -7.18875039757596-180.57352481571-186.952678101753-183.787706640935229.8272082174
#> outer loop: 17
#> -6.95384131780215-174.181080719961-187.936609954848-183.787706640935229.166483106556
#> outer loop: 18
#> -6.9525789516857-167.867863635072-187.951397844668-183.787706640935225.286444727927
#> outer loop: 19
#> -6.95216017510315-162.048875569435-188.92116205436-183.787706640935224.955687887465
#> outer loop: 20
#> -6.66281256060524-156.069833249585-188.933127113343-183.787706640935220.170884311034
#> outer loop: 21
#> -6.66134343731587-149.82069962412-188.939071517304-183.787706640935216.080602321946
#> outer loop: 22
#> -6.66093339360905-144.246896869333-189.754160275349-183.787706640935215.850482567777
#> outer loop: 23
#> -6.66086283302329-138.575764431927-189.932392389976-183.787706640935211.543137145562
#> outer loop: 24
#> -6.28084700237812-133.122184409321-189.934233859611-183.787706640935207.171868505397
#> outer loop: 25
#> -6.2801026129219-128.230914707117-189.972071572491-183.787706640935205.127642537616
#> outer loop: 26
#> -6.2800548487799-124.113914743005-190.931620861176-183.787706640935204.665282860171
#> outer loop: 27
#> -6.28005455627573-119.549358666513-190.931718886745-183.787706640935200.786645640312
#> outer loop: 28
#> -6.28004822905403-114.559354499013-190.931942659093-183.787706640935196.690096572967
#> outer loop: 29
#> -5.72383608763809-110.106059836356-190.932662164725-183.787706640935192.565238538002
#> outer loop: 30
#> -5.72249390574909-105.795668651159-190.938868051272-183.787706640935190.305827413031
#> outer loop: 31
#> -5.7221945461323-103.942065874093-191.84293642393-183.787706640935193.1193554971
#> outer loop: 32
#> -5.72214756996271-102.529553102901-191.922115127596-183.787706640935191.930682169872
#> outer loop: 33
#> -5.72209392848756-100.876963462084-191.922124617926-183.787706640935190.444330777343
#> outer loop: 34
#> -5.72203291592173-98.9454678632905-191.922137576682-183.787706640935188.710763265193
#> outer loop: 35
#> -5.72196412955988-96.6909454710423-191.922155946282-183.787706640935186.69565641059
#> outer loop: 36
#> -5.72188746559879-94.0636667459246-191.922183378447-183.787706640935184.364028942299
#> outer loop: 37
#> -5.72180342278283-91.011257368003-191.922227498545-183.787706640935181.687107312713
#> outer loop: 38
#> -5.72171370688634-87.4901358482224-191.922306762593-183.787706640935178.66184373861
#> outer loop: 39
#> -5.72162247854409-83.5045661151612-191.922476698454-183.787706640935175.367802683629
#> outer loop: 40
#> -5.72153882373598-79.2417902729959-191.922974840712-183.787706640935172.13876119915
#> outer loop: 41
#> -4.64685080936348-76.9691843723977-191.925801688354-183.787706640935169.3987400938
#> outer loop: 42
#> -4.64636241153534-75.3372755662499-192.255221098821-183.787706640935172.246640265296
#> outer loop: 43
#> -4.64635867491652-75.3373637985576-192.922015558643-183.787706640935173.321986309963
#> outer loop: 44
#> -4.64635867461363-75.3373646343763-192.922015565081-183.787706640935173.321928795621
#> outer loop: 45
#> -4.64635867460407-75.3373646602517-192.922015565281-183.787706640935173.321926941795
summary(res)
#> Length Class Mode
#> posterior 5 -none- list
#> optimisation 3 -none- list
#> PCA_viz 1 ggplot2::ggplot object
#> ELBO_viz 1 ggplot2::ggplot object
#> Seed_used 1 -none- character
plot(res)